To be, or not to be, a drug target gene
I have a drug target gene (or gene list) and I want to do a preliminary investigation to learn more about my gene and gene products of interest. But where do I start? Which genomic, biological or bioinformatics resources should I use? There are so many databases available. Here are just a few to get you started, and a clue to the data you can find.
Start with the gene
For an initial look at your gene and its products, why not visit Ensembl. Enter a query for your gene, remember that you may be querying with the synonym of approved gene symbol (for example FANCS is a synonym of BRCA1), so the results may not be exactly what you were expecting to see! Once you have identified your gene, take a note of the approved symbol as this will help your further database queries. Also note that Ensembl have assigned identifiers for the gene (ENSG), transcript (ENST) and protein (ENSP), these identifiers will be used to map data within Ensembl and between Ensembl and other databases.

Ensembl Gene details
On the gene page, Ensembl will provide genomic data, including the overall gene structure (how many exons, size of the CDS, etc) number of transcripts and proteins. There are also links to the phenotypes associated to the gene and the SNPs within the gene, as well as a full table of variants. Orthologues, paralogs and protein family members are also provided to give an overview of the taxonomy and ancestral “development” of your gene.
Some of our customers have taken advantage of our expert knowledge of Ensembl and have a customised installation that allows them to visualise and search their genome data. If you are interested in eaglensembl then please contact info@eaglegenomics.com.
Transcripts and proteins
From the gene page there is a table of transcripts and their biotype (protein coding, processed transcript etc). For transcripts there is haplotype data; some SNPs tend to occur together and inform the protein generating multiple protein forms, these are haplotypes. The consequence of the amino acid change, the overall frequency of that haplotype in the 1000 Genomes group is also provided. Interestingly, the reference genome may not encode the most frequent protein form.
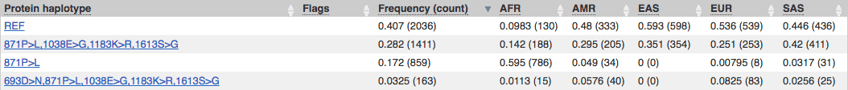
Ensembl Haplotypes
More detailed information about the protein can be found in UniProt. This can be used to supplement with protein architecture and and amino sequence level annotations, as well as provide links to many other resources that may be of interest to you.
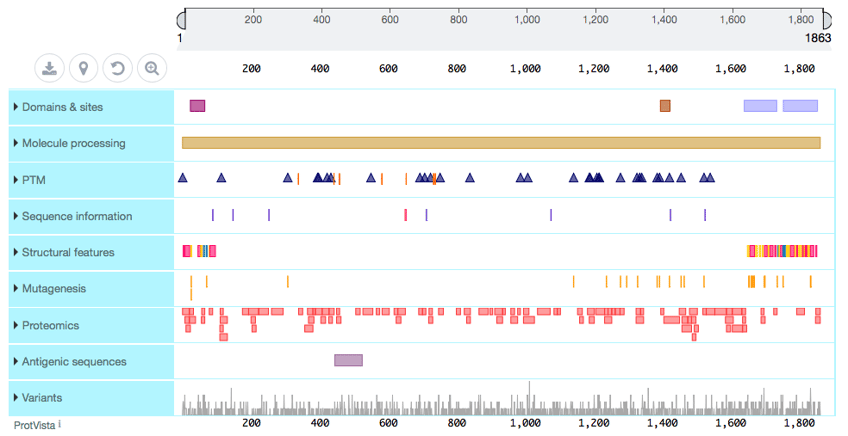 Protein features in UniProt
Protein features in UniProt
Sequence variations
We can augment the SNP information for each gene by also looking at the GWAS catalogue, GWAS Central, DbSNP, openSNP and OpenTargets. From all these resources a full picture can be developed about sequence variation and association with phenotypes and disease.
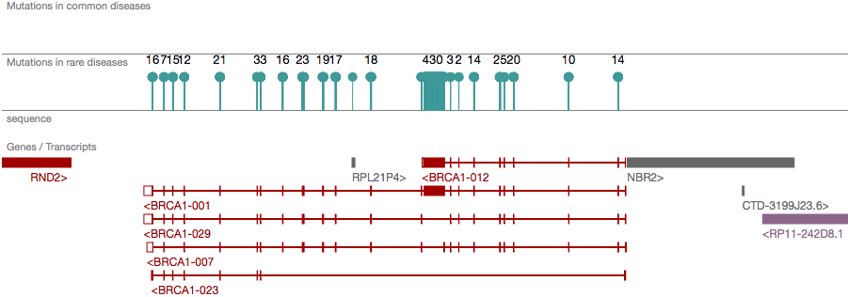 Gene vararions in OpenTargets
Gene vararions in OpenTargets
Association with disease
OpenTargets provides a search for a disease (and phenotype) and your target gene, the results include association score to summarise the strength of evidence for several attributes (somatic mutation, known drugs, pathways, RNA expression etc).
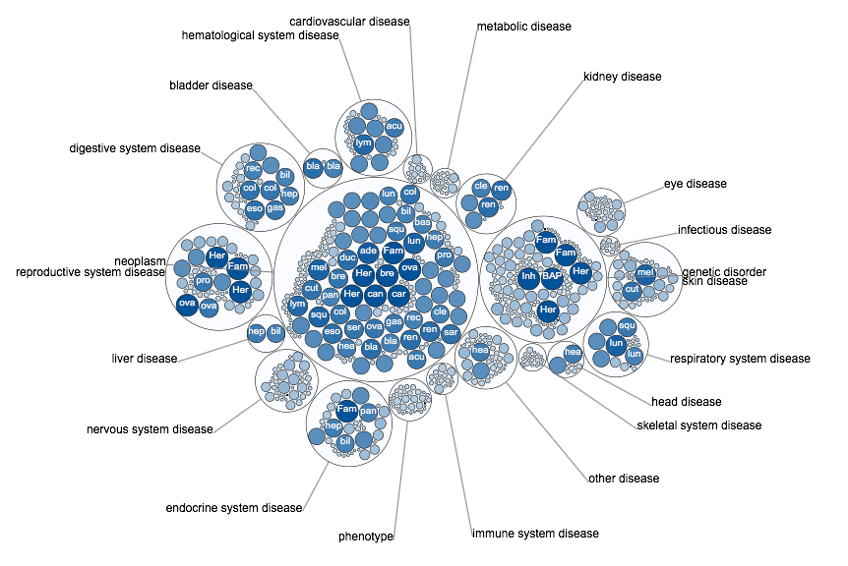 Disease associations in OpenTargets
Disease associations in OpenTargets
Tolerance to loss of function
The ExAC catalog can be used to calculate objective metrics of pathogenicity for sequence variants, and to identify genes subject to strong selection against various classes of mutation. A probability of loss of function intolerance is provided, if intolerance is low (near 0) this suggests humans are able to cope where both copies of your gene of interest are non-functional.
eQTLs
By analyzing global RNA expression within individual tissues and treating the expression levels of genes as quantitative traits, variations in gene expression that are highly correlated with genetic variation can be identified as expression quantitative trait loci, or eQTLs. The Genotype-Tissue Expression (GTEx) project provides a comprehensive identification of human eQTLs, providing a valuable basis on which to study the mechanism of that genes regulation.
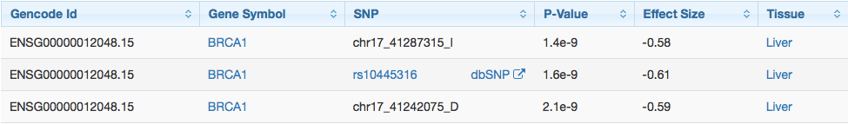 eQTLs from GTEx.
eQTLs from GTEx.
Clinical trials
ClinicalTrials.gov is a registry and results database of publicly and privately supported clinical studies of human participants conducted around the world. You can search for your gene of interest to learn if has been involved in any registered clinical trials.
Conclusion
Following these suggestions, in addition to reading publications, will provide you with an excellent overview of your gene of interest, its structure, products, variants and their functions. Each gene can have thousands of sequence variations and you will be able to determine those that have a known association to your disease or phenotype of interest. Knowing if your target gene can tolerate loss of function is a good indication of safety and links to clinical trials will let you know if others are also interested in the same gene. Sounds like a good start to me!
May 2017 will see the first iteration of a new product, eaglecurate. The platform will provide functionality to link and automatically annotate data sources or specific R&D data with the appropriate semantic content which could reduce the time taken to manually investigate genes of interest. Watch this space!
If you would like to discuss how eaglecurate and eaglensembl can help further your pursuit for genomic, biological or bioinformatic research, get in touch on +44 (0) 1223 654481 or drop us a line with our specialised form.
